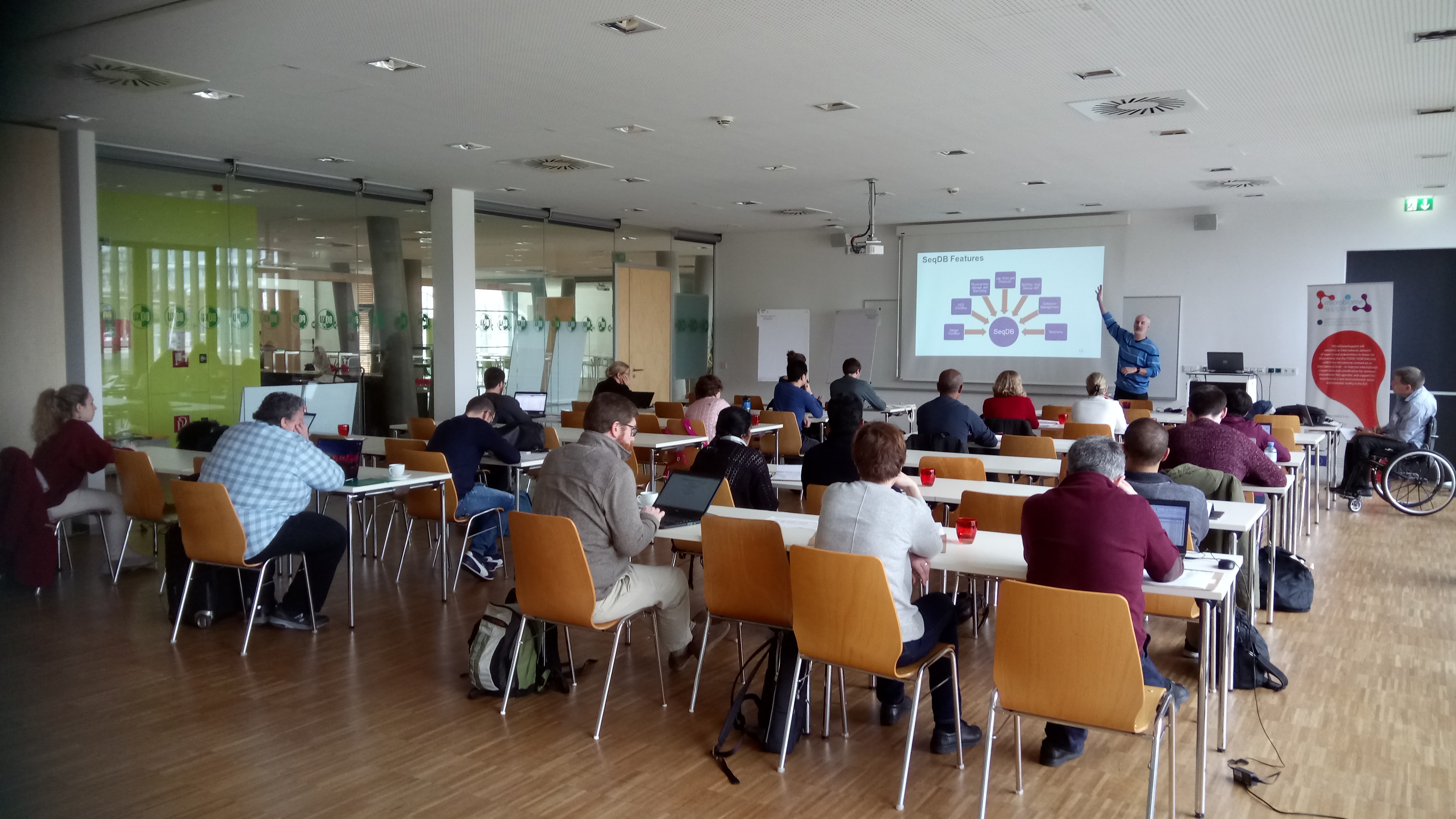
Around 25 partners from the MicrobiomeSupport consortium joined a workshop on 7 March 2019 to discuss what microbiome researchers need from biobanks and data storage. After setting the scene with introductory presentations on the status quo of culture collections, biobanks and how microbiome research data is stored and analysed, the scientists discussed the limitations of current systems and specific needs within the microbiome research field in groups. The consortium partners convened to agree on writing a couple of white papers that outline recommendations and guidance of microbiome research needs from biobanks and data storage.
MicrobiomeSupport partners and advisory group members discussing what we need from biobanks and data storage for microbiome research.
What are biobanks and why are they important?
Biobanks store biological material (e.g. blood, soil, water, micro-organism samples) for use in research. They are crucial for researchers:
- to show that repeated experiments result in the same outcome
- to maintain high research quality
- to protect IP, assests and investments
- to conserve biological material (for instance, for habitat reconstruction)
- for knowledge and information
- for insurance for products released into the environment
Why is data storage important?
Data banks store the data and results associated with biological samples. In microbiome research data a lot of server space is needed to store these data. Data include:
- The different organisms that are in a microbiome sample
- DNA sequence data (genomic data)
- Functional data (e.g. what products micro-organisms produce and how they work, often which effect they have on their environment)
Further, researchers recognise the importance of meta-data more and more to ensure that experiments can be repeated successfully and there is transparency in the scientific approach. Meta-data include:
- The researchers, time and location where samples where taken from
- How samples were stored, transported and analysed
- How data were stored and analysed
- Which protocols were used
- Who has done what to ensure the right people receive credit
What do biobanks and data storage have to do with MicrobiomeSupport?
Within MicrobiomeSupport, scientists from all disciplines are trying to come up with standardised research protocols or quality standards to maintain in microbiome research to:
- avoid having to repeat experiments to answer the same question
- maintain research ethics
- ensure that we can compare individual studies better and avoid having seemingly contradictory data
So, what has been discussed in the workshop?
The consortium partners considered what biobanks, resource collections and data storage systems currently exist in different scientific disciplines, their benefits and limitations.
For instance, did you know that microbiome research is still very limited by computing power?
It soon became clear, that for microbiome research currently existing bio banks are not sufficiently equipped to deal with microbiome samples. Further, data storage systems do not yet exist in such a way that data can be collected in a harmonised way or that data are assessed for their quality. For neither microbiome samples or data storage there is a clear responsibility on whom is responsible for providing the relevant infrastructure for storage.
What now?
The consortium partners will now assess best practice examples, see what can be used or adapted for microbiome research, and foster alignment with currently existing biobanks, resource collections and data storage system administrators to ensure that standards in microbiome research can be strengthened. The aim will be to have two white papers later on this year to outline specific suggestions, provide guidance and suggestions on how the current limitations in microbiome research can be solved.
Want to be informed when this is going to happen? Sign up for our newsletter here.